Publications
De novo protein design by citizen scientists (PDF)
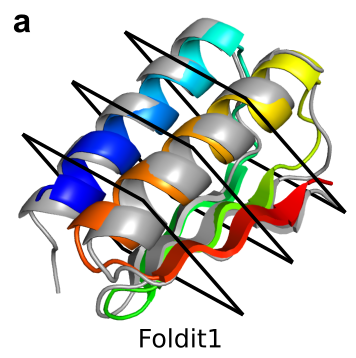
Proteins designed by Foldit players fold accurately, with high stability, and span a diversity of protein folds.
Koepnick B, Flatten J, Husain T, Ford A, Silva D, Bick MJ, Bauer A, Liu G, Ishida Y, Boykov A, Estep RD, Kleinfelter S, Nørgård-Solano T, Wei L, Foldit Players, Montelione GT, DiMaio F, Popović Z, Khatib F, Cooper S, Baker D. De novo protein design by citizen scientists. Nature (2019)
Determining crystal structures through crowdsourcing and coursework (PDF)
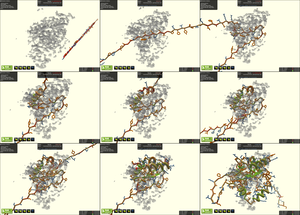
Foldit players build a high-quality protein model, into electron density from experimentally-phased crystallography data.
Horowitz S*, Koepnick B*, Martin R*, Tymieniecki A, Winburn AA, Cooper S, Flatten J, Rogawski DS, Koropatkin NM, Hailu TT, Jain N, Koldewey P, Ahlstrom LS, Chapman MR, Sikkema AP, Skiba MA, Maloney FP, Beinlich FR, Foldit Players, University of Michigan students, Popović Z, Baker D, Khatib F, Bardwell JC. Determining crystal structures through crowdsourcing and coursework. Nat. Commun. 7 (2016)
Effect of Substitution on the Optical Properties and HOMO−LUMO Gap of Oligomeric Paraphenylenes (PDF)
Quantum mechanics calculations and absorption spectra of a para-phenylene dimer with various chemical substitutions.
Koepnick B, Lipscomb JS, Taylor DK. Effect of Substitution on the Optical Properties and HOMO−LUMO Gap of Oligomeric Paraphenylenes. J. Phys. Chem. A 114 13228–13233 (2010)
